Global proteome quantification: dreams or reality?
Brownridge, P., Holma, S.W., Gaskell, S.J., Grant, C.<., Harman, V.M., Hubbard, S.J., Lanthaler, K., Lawless, C., Ocualain, R., Sims, R., Watkins, R & Beynon, R.J. (2011) Global absolute quantification of a proteome: Challenges in the deployment of a QconCAT strategy. Proteomics, (in press)
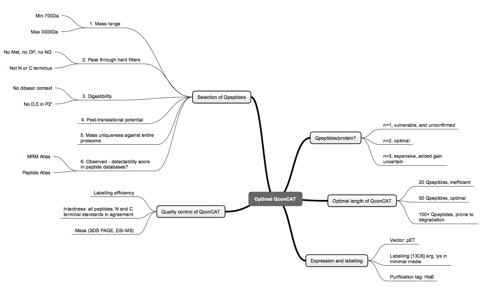
In this paper, we discuss the challenge of large scale quantification of a proteome, referring to our programme that aims to define the absolute quantity, in copies per cell, of at least 4,000 proteins in the yeast Saccharomyces cerevisiae. We have based our strategy on the well established method of stable isotope dilution, generating isotopically labeled peptides using QconCAT technology, in which artificial genes, encoding concatenations of tryptic fragments as surrogate quantification standards, are designed, synthesised de novo and expressed in bacteria using stable isotopically enriched media. A known quantity of QconCAT is then co-digested with analyte proteins and the heavy:light isotopologues are analysed by mass spectrometry to yield absolute quantification. This workflow brings issues of optimal selection of quantotypic peptides, their assembly into QconCATs, expression, purification and deployment.